On my journey to becoming a cheminformatician, it took me a little time to realize philosophically what it truly meant to me. Perhaps this PhD is more introspective than I thought.
It wasn’t about hunting drugs, fingerprints recognition, machine learning, organic chemistry, code, devops, graph theory, or pretty visuals. That’s the easy stuff. Rather, it was communication.
Scientists struggle with translating their ideas to the common public and even more so they struggle to talk to each other. We are very involved with our work, we forgot to listen to others. Here’s one of the problems I’ve noticed, which you can call as “knowledge” gap but rather it’s a “communication” gap.
The organic chemist has, naturally, a tough time talking to a physicist. Because we disagree. That’s not ego but a challenge to face as we start linking data. One of the major reasons of this disagreement is the charge of a molecule. We as organic chemists are trained on functional groups and use Nuclear Magnetic Resonance (NMR) to justify electronegativity. Physicists are more mathematical based then the organic chemist (we work on color and movement) and use harmonic vibrational frequencies to validate the movement of a molecule. With energy equations ruling the perceived potential energy and number parameters for each type of molecule in scenario then can predict some geometry and in turn use that to generate charges.
For example, the General Amber ForceField (GAFF) and Charmm General Forcefield (CGenFF) use to different sets of equations and guess the series of green dots until the numbers line up with their predict quantum mechanics of what the charge will be.
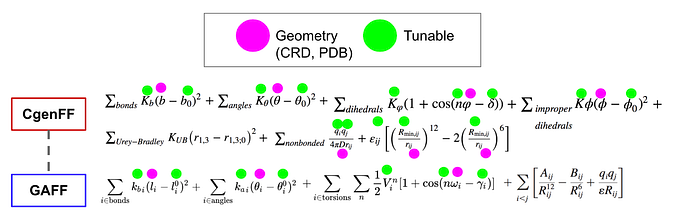
So the organic chemists have a feeling, a intuition, where the physcists are more methodical. Or that’s what I used to think.
In the cheminformatic community, around the 80s the David et al came up with a language easy for the medicinal chemists to write alphanumeric characters into a 1–D string which also contained the rough geometry representation of a molecule:
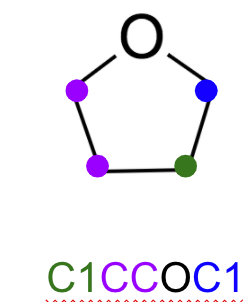
This was easy for me to read and translate yet there is no charge information store within this string. I have to intepret that myself (that’s what I was paid to do in my previous life).
Simiarly, the physicists designed their own language to explain the molecule by categorizing one level deeper: the atom types. In my example the CGenFF Atom Type Engine.

This language is a little more complex to understand but also explains the same concept at it’s core using the same philosophy as SMILES. This was remarkable to me, a convergence has happened, and from what it seems like independently of each other. This type of phenomena I have seen once in my life before with nuclear warhead zipper chemistry.
Both languages actually have benefits but store the same information from different sources. Yet the values disagree so the two communities disagree. And what is worse there are several demoniations of the same concept applied with different terms so the numbers disagree even moreso (GAFF, CGenFF, OPLS, SMIRNOFF). It’s actually created a heated rivalry amongst forcefield developers, myself of which I am now included as being part of one of the sides.
The Atom Type language was used to help the physcists talk to each other as well as SMILES has done for the organic chemistry community like IUPAC. Recently, I have read papers trying to circumvent the atom type language or blantantly ignore it: work from Mobley et al.

While reading this paper, although science, philosophically I felt it was immoral. Using the word Escape is a slap in the face to the physicist community. Rather than trying to understand the Atom Type Language they chose to actively not understand it and learn how to use it futhermore — it just looks like it adds more complexity.
This is actively not what a cheminformatician would do. We like languages, we like to connect, and not to reinvent to call it our own. It’s not our place to do so. Atom Type language contains the physicists values that correspond the thermodynamic properties of a system. SMILES does not contain that information but rather the perception of what molecule will look like. It is easy for us medicinal chemists to understand and the Atom Type Language is easy for physicist to understand. The best course of action I believe would be is to link them.
Please wait for part 2 and I will explain the output of CGenFF in greater detail to ameliorate the understanding of what is going on in the world.
